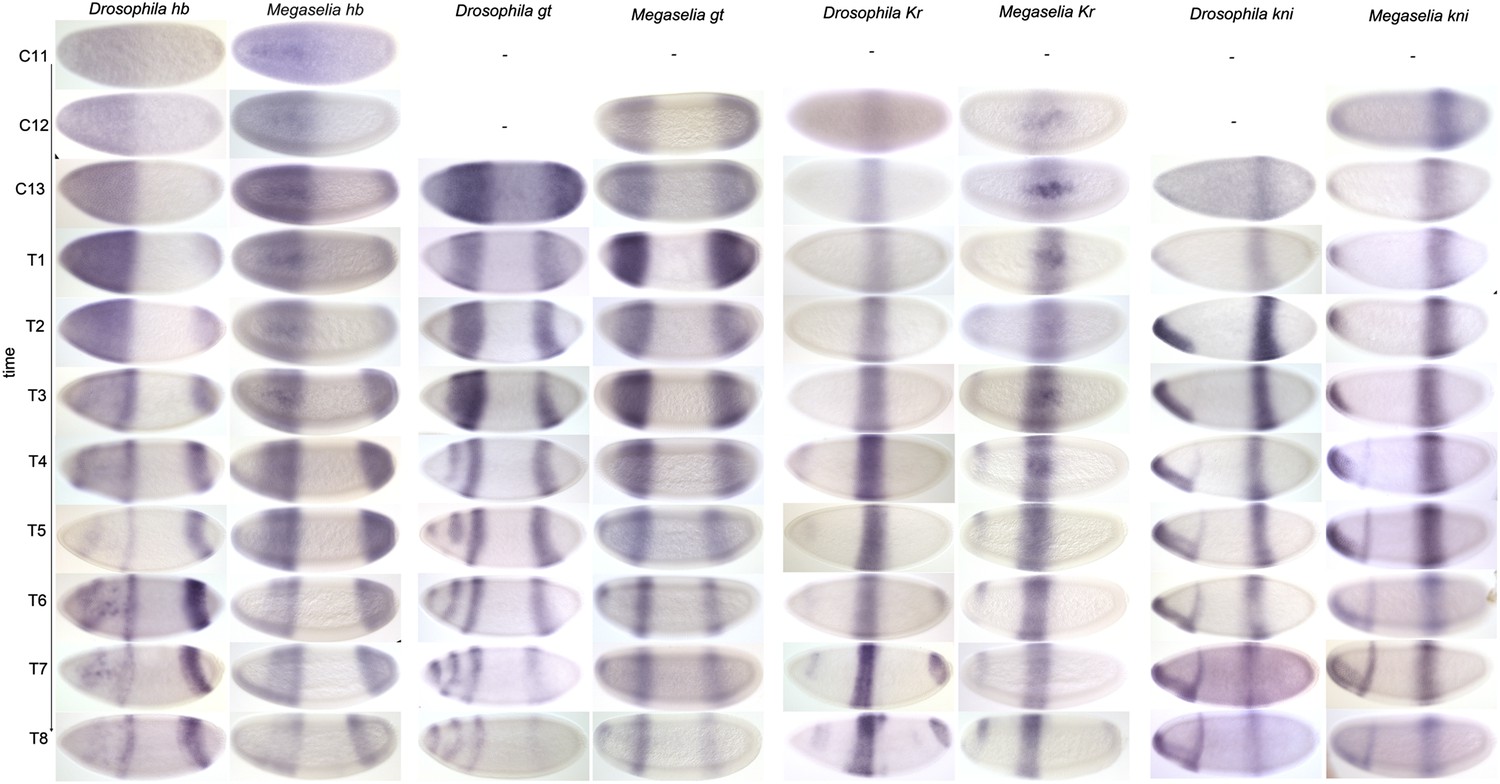
PLOS Biology: A damped oscillator imposes temporal order on posterior gap gene expression in Drosophila

Whole-Embryo Modeling of Early Segmentation in Drosophila Identifies Robust and Fragile Expression Domains: Biophysical Journal

Dynamic Maternal Gradients Control Timing and Shift-Rates for Drosophila Gap Gene Expression | bioRxiv

Drosophila Neuroblasts Sequentially Express Transcription Factors which Specify the Temporal Identity of Their Neuronal Progeny: Cell

Dynamic Maternal Gradients Control Timing and Shift-Rates for Drosophila Gap Gene Expression | bioRxiv
PLOS Computational Biology: Reverse-Engineering Post-Transcriptional Regulation of Gap Genes in Drosophila melanogaster

Comparison of larval and adult Drosophila. Drosophila segmentation genes Gap: Kruppel (Kr),knirps (kni), hunchback (hb), giant (gt),tailless tll), Pair. - ppt download

Decoding in a genetic patterning network. a. In the early Drosophila... | Download Scientific Diagram

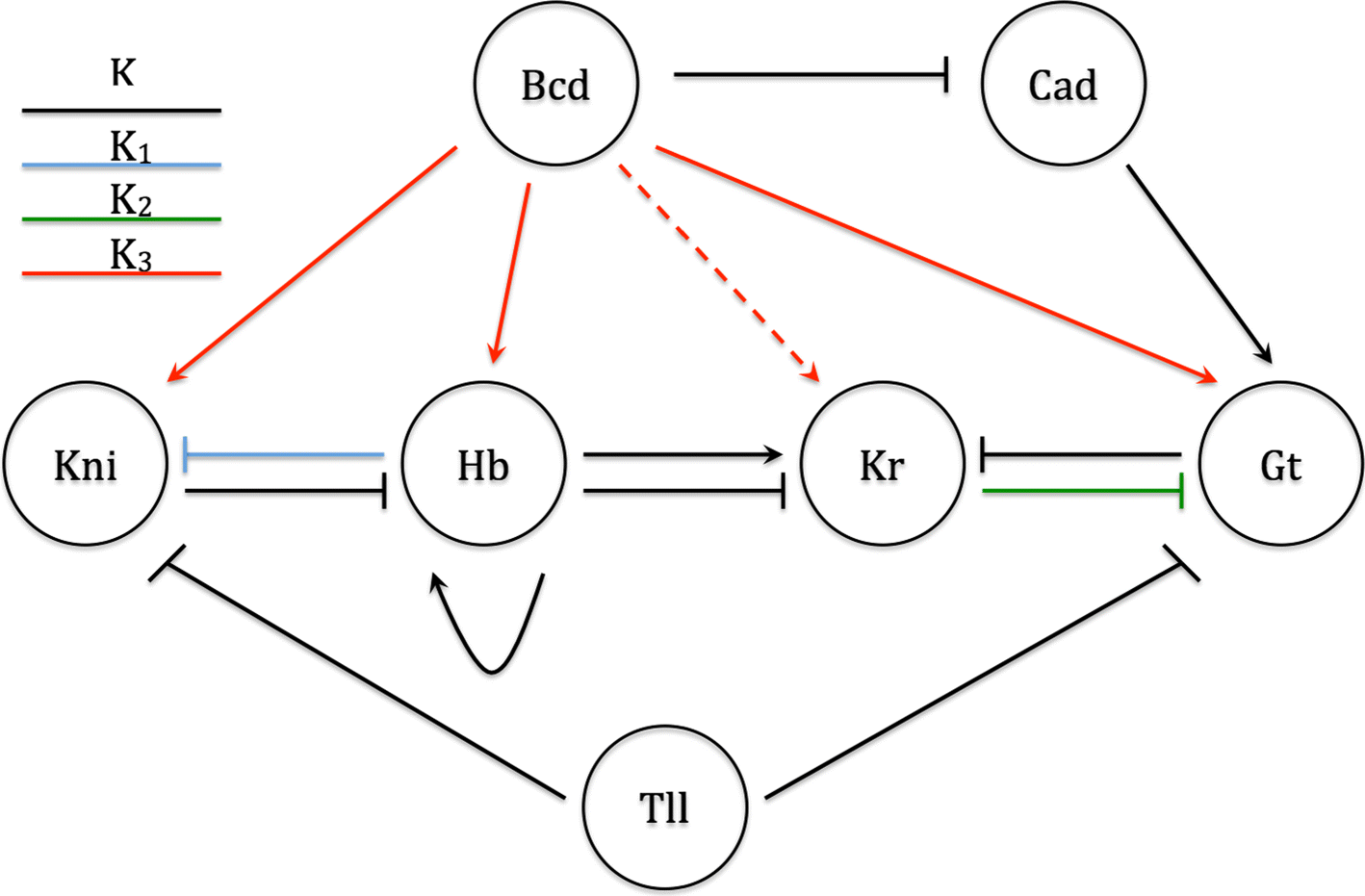
Dynamical Analysis of Regulatory Interactions in the Gap Gene System of Drosophila melanogaster | Genetics

![PDF] A logical analysis of the Drosophila gap-gene system. | Semantic Scholar PDF] A logical analysis of the Drosophila gap-gene system. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2217795bf54883bd97e6cbfc85fa4bb7b2a6e17e/5-Figure2-1.png)